Credit for featured photo: Caroline Donahue from Flicker available via Creative Commons (CC).
Whether it is blood splattered at the scene of the crime, semen swabbed from sheets in a hotel room, or even saliva collected from a bite mark, body fluids have a heavy presence in forensic science. This type of evidence acts as a source of DNA and provides strong evidence to support or refute a claim in court. Thus, body fluid identification is a crucial part of the forensic field.
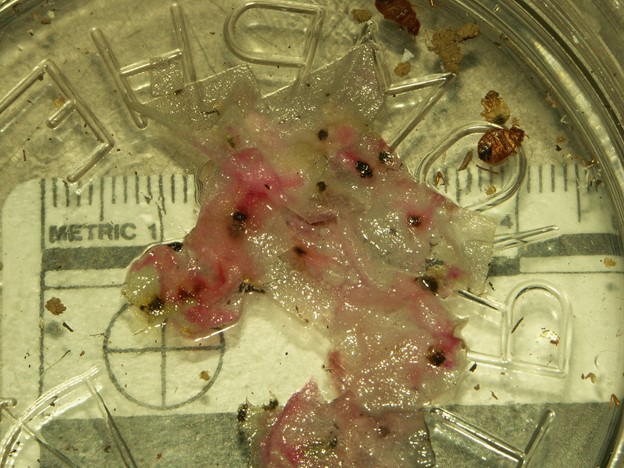
Traditionally, color tests such as the Kastle Meyers Test, which tests for the presence of blood (Figure 1), and the Acid Phosphatase Test, which tests for the presence of semen, are used to determine the likely identity of body fluids (presumptive identification). The color tests chemicals (like those used in the examples previously mentioned), may cause a color change when added to the stains and indicate the presence of a specific bodily fluid. Despite their usefulness for presumptive identification, these color tests are insufficient for confirmation of bodily fluid identity due to their false positive and false negative rate (incorrectly indicating a positive when the result is actually negative and vice versa).
Liquid chromatography (LC) is a common method used to confirm the identity of body fluids. In LC, a liquid carries the sample through a column coted in a solid gel. The sample separates based on whether its components have a greater attraction to the liquid (thus separating first) or the gel (thus separating last). Based on the specific biomarkers present in body fluids (such as hemoglobin for blood), scientists identify the body fluid. While this method is reliable, the sample preparation and pretreatment, choosing and optimizing of the LC method parameters, and data analysis associated with it is time consuming. Thus, alternative methods for the identification of body fluids are of interest.
One such method is matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS). In MALDI-MS, much of the MS procedure is the same, except for the ionization process. Instead of using a stream of electrons to ionize a sample like in simpler MS methods, an analyst must first disperse the sample in a matrix solution. Then, they irradiate the matrix with a laser, causing it to absorb energy, vaporize along with the sample, and transfer a proton to the sample, thus ionizing it. Then, the ions are accelerated by an electric field toward the detector which plots out the intensity of the ions’ signal vs their mass-to-charge ratio (m/z).
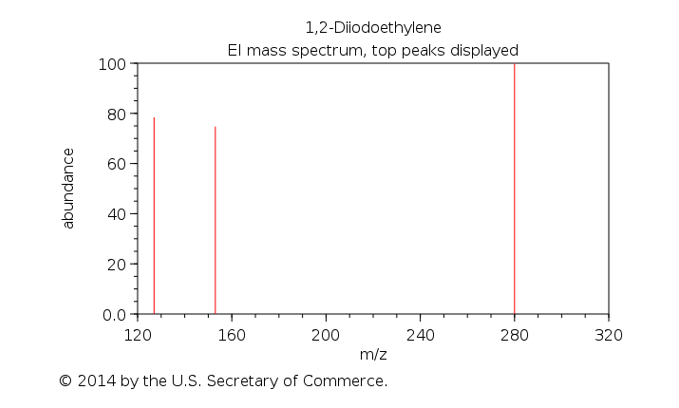
This tool emerged as a crucial tool in biology and chemistry due to its high resolution and speed. Zongxiu Nie and his colleagues investigated if this method could be used to identify five forensically relevant body fluids (blood, saliva, sweat, semen, and urine) without any pretreatment.
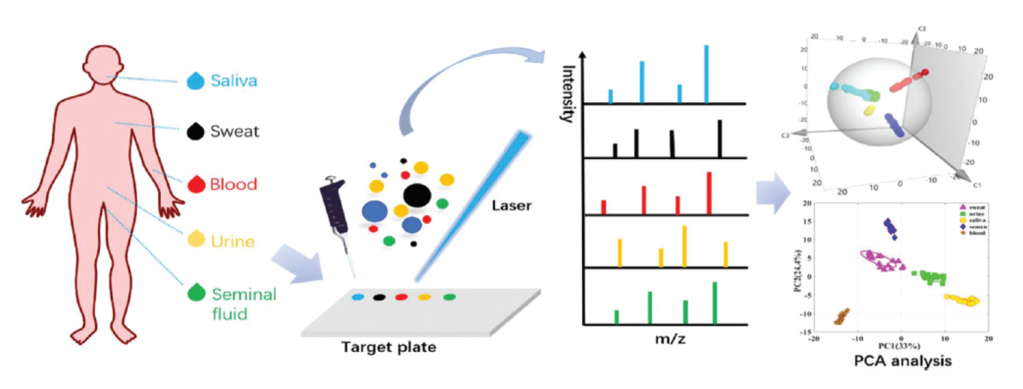
The general scheme for the procedure is shown below in Figure 3. First, the researchers collected blood, saliva, sweat, semen, and urine from nine volunteers and diluted them to the appropriate concentration before mixing them with the matrix solution, 1-(naphthyl) ethylenediamine dihydrochloride. They stored these solutions for various lengths of time at -20 °C to evaluate the effect of refrigeration time on the results. Zongxiu Nie and his colleagues also tested body fluid samples that they deposited on cotton, fabric, and tissue before dissolving them in distilled water and combining them with the matrix solution. They recorded the MALDI MS data for these samples and preprocessed the resulting data. The scientists then used machine learning models to differentiate between the five body fluids. Finally, they tested these models using ten unknown body fluid samples that had been deposited on different materials (cotton, fabric, or tissue).
The researchers found that MALDI MS detected biomarkers associated with the analyzed body fluids. Further, they found that storage time of the body fluids, and deposition on different materials appeared to have no effect on the ability of the method to detect the biomarkers.
The scientists performed statistical analysis using a machine learning model known as Principle Component Analysis (PCA). This method plots data points based on different scores in a three-dimensional plot which results in different groups within the data set being visually separated into distinct clusters. The PCA results showed that the five body fluids could be clearly separated into five different groups (Figure 4).
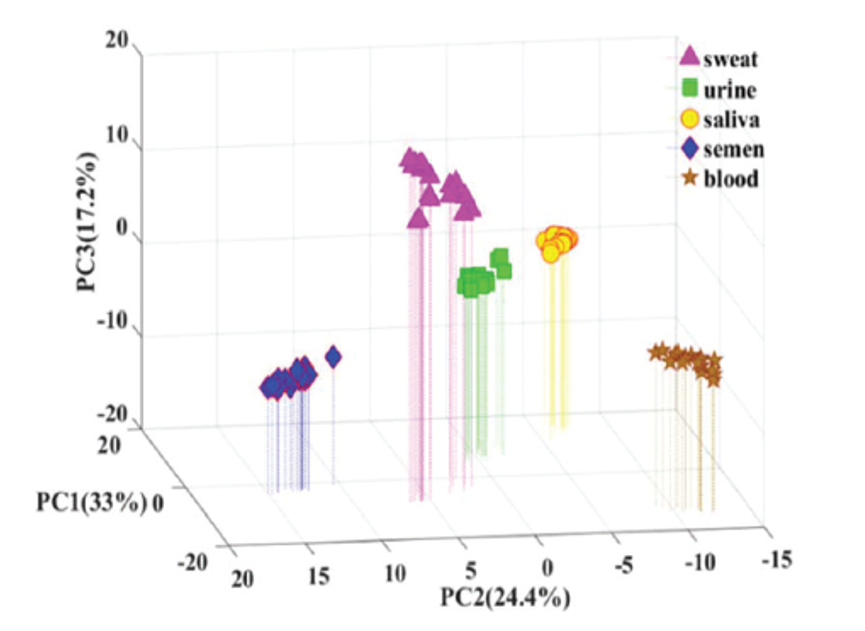
When Zongxiu Nie and his colleagues tested the ten unknown samples using the PCA model, nine out of ten were categorized correctly. They couldn’t classify one saliva sample using PCA, but by looking at the unique MALDI MS peaks present in its mass spectrum, they were able to correctly identify it as saliva.
This MALDI MS technique coupled with the machine learning model shows a lot of promise, as the researchers have demonstrated that it can differentiate between five forensically relevant body fluids. Traditional methods often test for one body fluid at a time, can be prone to false positive and negative results, and may require pretreatment. This method tests for five body fluids without pretreatment while maintaining a high sensitivity and accuracy.
Such a method is crucial in forensic science where body fluid evidence makes a regular appearance and is of great importance. However, the disadvantage still remains that like LC, this method is destructive, meaning it consumes the analyzed sample. This is not ideal in forensics where it is best to preserve as much of the original sample as possible. Thus, Zongxiu Nie and his colleagues are considering looking into another form of ionization, Desorption Electrospray Ionization (DESI), which may allow for the non-destructive analysis of the five body fluids.
| Title | Direct Identification of Forensic Body Fluids by MALDI-MS |
| Authors | Yuming Jiang, Jie Sun, Xi Huang, Huixia Shi, Caiqiao Xiong, Zongxiu Nie |
| Journal | Analyst |
| Year | 2019 |
| Link | https://doi.org/10.1039/C9AN01385G |
| Citation | Y. Jiang; J. Sun; X. Huang; H. Shi; C. Xiong; Z. Nie; Direct identification of forensic body fluids by MALDI-MS. Analyst, 2019, 144, 7017. DOI: 10.1039/C9AN01385G |
