Credit for Source Photo: Gum by Pxfuel
The human body provides an enormous amount of genetic information, from our blood and hair, to the DNA of the microbes that live in and on our bodies, known as the human microbiome. We classify the different microbiomes in our body based on the body site they inhabit, such as the skin or stomach. Forensic scientists have previously studied microbiomes found in the environment to reveal forensically relevant information, such as geographic location or ethnic population.
Highlighted as one of the best scicomm pieces of the week in February 2021 by ScienceSeeker, a large-scale science news aggregator, alongside articles from Scientific American and Massive Science.
Alba Leila Satari, Àngela Vidal-Verdu Guillén, and Manuel Porcar from the Institute of Integrative Systems Biology, a joint research center between the University of Valencia and the CSIC, studied the microbiome of the mouth, aka the oral microbiome, to identify the different bacteria that develop on chewing gum. They discovered that the collective bacteria present on chewed gum evolve into their own environmental bacteriome, useful in forensics to shed light on whether someone was present in a particular location or for uncovering hidden graves.
Forensic scientists can already extract the “chewers” DNA from discarded gum and obtain an individual DNA profile. But trapped in the sticky residue is also a fraction of the oral microbiome, potential toxins (such as drugs or poison) and pathogens (like viruses) residing in our mouths. Scientists can use these characteristics to identify geographic regions of the gum consumer and even their ethnic background, placing a potential suspect at the location of a crime or narrowing down a suspect list. The survival and succession of these bacterial environments over time has received only limited research attention. Satari and her team shine a light on how the bacteriome of chewing gum from five different countries changes once discarded from our mouths and placed on an outside surface.
To identify the bacteria present, scientist rely on taxonomy, or Latin-based naming system that encompasses eight categories, from the overarching Domain Bacteria down to the individual Species. A genetic sequencing technique called 16S rRNA sequencing can identify all of these names down to the 7th Genus level for bacteria, which is the last step before Species. This well-used method is bacteria specific and distinguishes between genera by small differences on their ribosomal RNA (rRNA).
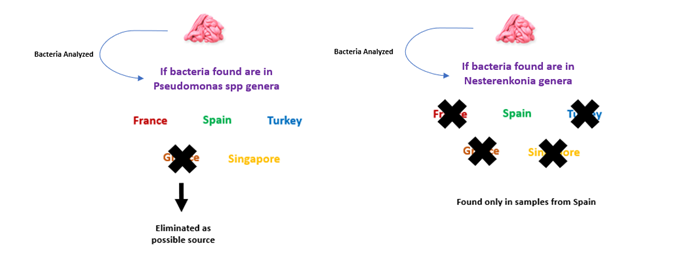
The team sourced the bacteria in their gum samples from Spain, France, Singapore, Turkey, and Greece, and found that although the samples showed important variation among locations, several bacterial genera were common amongst the different countries but in different abundance. For example, the genus Kocuria was the most abundant genera in samples from Singapore and France and found about 50% of the time, whereas in Greek samples, the most frequent genera was Deinococcus with a 25.2% frequency, followed by Kocuria with a relative abundance between 10 and 16%. The scientists even showed that if you found Pseudomonas bacteria in the sample, that eliminated Greece as the origin, whereas if you found Nesterenkonia, you knew it could only be from Spain out of the five countries (Fig 1). Coupling these factors together greatly narrowed down the geographic origins of the sample.
In this work, Satari and her team characterized all the bacteria present aka the bacteriome on a piece of discarded chewing gum. There is moderate difference of the kinds of bacteria present among the samples based on geographic location and population group. This is especially interesting as previous studies have found that although our genetics influence the bacteria in our oral microbiome, environment has a greater influence. The scientists looked at geographic regions separated by large stretches of land, however, in theory, this same system could also work on a smaller scale, such as within a single state or even city, if enough diversity can be found. If a crime is committed, and a piece of gum is found at the crime scene, the bacteriome could be used to identify which ethnic population the bacteria is most found in (e.g. Asian, European, African, Hispanic populations, etc.). This is especially useful when only a partial DNA profile can be generated, or no profile at all by narrowing down the suspect list.
While normally when discovering chewed gum stuck to your chair, we respond with disgust, CSI techs and lab scientists find the sample another treasure trove of forensically relevant information.
| Title | The wasted chewing gum bacteriome |
| Authors | Leila Satari, Alba Guillén, Àngela Vidal-Verdú & Manuel Porcar |
| Journal | Scientific Reports |
| Publisher | Nature |
| Year | 2020 |
| Link | https://www.nature.com/articles/s41598-020-73913-4#citeas |
