After a whole age of Arda’s of absence, the One True King returns to the throne of Elendil and Isisldur. On the year 3019 of the third age, Aragorn, son of Arathorn is crowned king of Gondor under the name of Elessar. His legitimacy to the throne is based on an un-interrupted paternal line which began more than 3000 years before the events of the J.R.R. Tolkien’s The Fellowship of the Ring.
But is Aragorn’s claim truly valid, that he is a direct descendant of King Elendil? If we assume yes and estimate an average 25 years per male in each generation, there would exist 120 generations between Isildur and Aragorn. However, Aragorn is not just a human, but a resident of Tolkein’s imaginary world of Arda and the land of Numer. As a Numenorean, their increased average lifespans are closer to 75 years per male generation, resulting in a 40-generation separation between those two characters.
The first part of our Royal Ancestry series by guest author Dr. Maria Sonia Fondevila dives into the determination of long-distance kinship by evaluating the ancestry of one of J.R.R. Tolkien’s beloved and most important protagonists; Aragorn II, son of Arathorn II and Gilraen, 16th Chieftain of the Dúnadan of the North, and the heir of Isildur Elendil’s son of Gondor.

Humans are diploid organisms, where half of our chromosomes come from each of our parents, of which theirs originate from the random mixture of their parent’s chromosomes, and so one and so on. We share an increasingly smaller portion of our genome with our remote ancestors as we travel further back in our lineage. Theoretically, if we could track that portion of DNA shared with a remote ancestor and find it in our own DNA, we can establish a proven relation – but like most things, tracking a single spot in our genetic code is not simple.
In order to establish any potential kinship, forensic geneticists analyze many DNA polymorphisms and compare them to the remains of the alleged ancestor or to those of a suspected living relative. If these individuals are related, called identity by ancestry (IBA), they would share several of these polymorphisms. Complicating this analysis, however, are the number of different types of polymorphisms within the population´s limited genepool and their natural frequency. There is always the possibility that two people may share a specific combo of variants just by random chance, called identity by state (IBS).
The key to distinguishing between IBA and IBS is to calculate the probability of both situations. IBA can be calculated by multiplying the shared portion of the genome between the living relative and ancestor plus complicating factors such as polymorphism mutation rates, inbreeding and random chance. To determine IBS, just calculate the random chance portion. Forensic geneticists then compare the long-distance kinship probability through Bayesian calculation and generate a likelihood ratio. Likelihood ratios tell us how likely one hypothesis is over the other, such as if the suspect’s blood on the deadly weapon supports the prosecution’s theory of the defense’s theory.
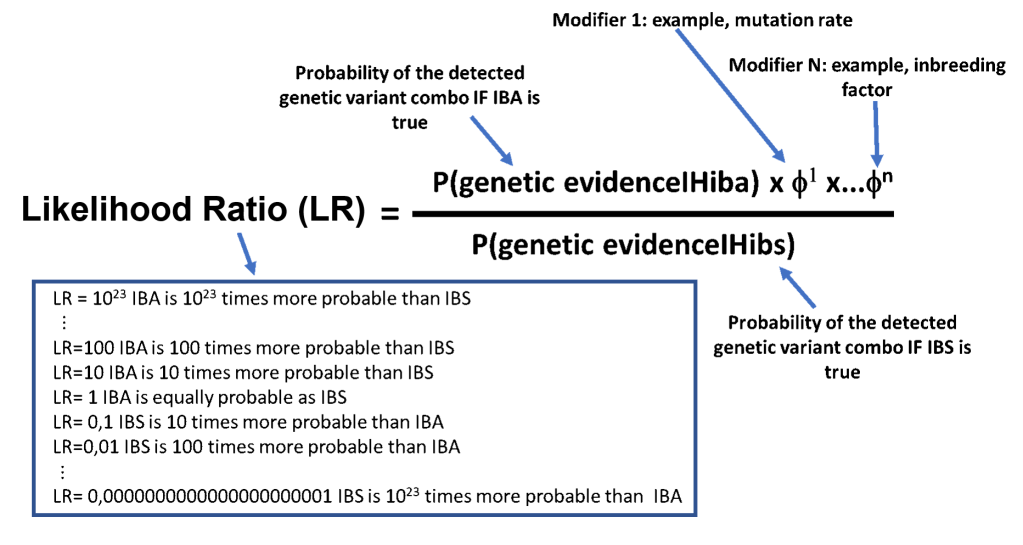
Thus, the smaller the probability of IBS, the greater chance that the likelihood ratio will be in favor of IBA, or true long-distance kinship. However, several issues factors significantly complicate the calculation of the likelihood ratio.
- The shared portion of the genome reduces more and more as the separation between the person in question (Aragorn) and the ancestor (King Elendil) grows larger. Finding that diminishing genetic spot when we search for polymorphisms may become impossible – an extreme version of looking for a needle in a haystack. Hence, the analysis loses effectiveness as the separation between an individual and their ancestor grows larger.
- Inbreeding greatly decreases genetic variability in populations, which mean shared polymorphisms will appear more often in a lineage. However, because these polymorphisms appear more often in the population, they become a higher frequency and therefore support the probability of both IBA and IBS.
- There is always a chance that the shared polymorphism combo between two related individuals is partly or totally random. The greater the separation between an individual and their ancestor, the larger the probability of random chance grows.
- Evolution drives the frequency of polymorphisms, and some will be naturally higher or lower in a given population. A high-frequency polymorphism, i.e. one that is more common in a population, gives a greater chance of IBS; a low frequency lowers this probability. The presence of rare polymorphism significantly helps with long distance kinships but finding higher frequency polymorphisms is much likelier.
Knowing all this, let’s revisit our original problem, whether we can determine that Aragorn is Isildur´s heir or not. Based on our list of complicating factors, including the 40 generations between Aragorn and Isildur and the inbred northern Dúnedain population, our final hypothesis remains inconclusive. Sadly, our current genetic tracing only accurately follows 6 generations back, which is way too small to determine Aragorn´s kin to Isildur. As far as we can analyze nowadays, Aragorn, son of Arathorn, Captain of the northern Dúnedain would be as related to his alleged remote ancestor as they would be to the Prancing Pony´s Innkeeper.

Guest author Dr. Maria Sonia Fondevila graduated in 2001 with a B.S. in biology at the University of Santiago de Compostela (USC) in north-western Spain. She started her graduate studies at the Institute of Forensic Sciences within the same university, specializing in forensic genetics and achieving her PhD in 2009. Afterwards, she worked as a post-doctoral researcher at the National Institute of Standards and Technology in Maryland. She then returned to USC and combined her research in the forensic genetics field with casework investigation. Her main research focuses on single nucleotide polymorphism (SNP) sequencing and InDel application to criminalistics, biogeographical ancestry and external visible characteristics prediction, highly degraded DNA analysis and study of DNA postmortem degradation. Currently, she coordinates the group of Human Identification of the USC spin-off genetic analysis company Genome4Care.
