Feature photo by Gustavo Fring from Pexels
On a frigid early morning in Karlstad, Sweden, CSI Brigitte Jansson arrived at the scene of a reported rape. Next to the outline of the victim’s body were splatches of blood – but amongst the white of the snow in the dark of dawn, no semen was visible. Jansson took her ultra-high-powered UV light and turned its rays onto the snow. There amongst the pristine fluffy powder were several spots of frozen semen. Laboratory technicians in Jansson’s district compared the sperm present in the frozen sample to the sperm on the victim and found a probabilistic match. This was essential as a full DNA profile could not be identified from the semen sample on the victim. The DNA profile from the frozen semen linked the suspect to the victim, and the rapist was convicted.
Identifying what body fluid in addition to whom the fluid originated from is crucial for forensic investigations, notably, to connect suspects with criminal acts, as shown in the aforementioned example. However, semen is not the only fluorescent body fluid, let alone the only fluorescent liquid – urine, sweat, saliva (and even antifreeze and bleach) all fluoresce strongly under UV light. To definitively identify the type of body fluid, laboratory technicians perform serological tests such as microscopic examination (looking for the sperm in semen) or chemical tests (like the Takayam Crystal test which forms red, feathery crystals in the presence of the iron in blood’s heme protein). These tests, unfortunately, require large amounts of sample as their minimum input – sometimes up to several drops of liquid, which can be difficult with dried semen or saliva – and do not function properly with degraded samples (such as bleach-cleaned latent blood).
That’s what motivated Dr. Keiji Tamaki, MD and Professor of Forensic Medicine at Kyoto University and lead investigator for the study, to investigate the potential of microRNAs (miRNAs) to identify body fluids [Fig 1]. These hardy RNA strands, averaging approx 22 nucleotides in length, are extremely resistant to external damage, making them ideal for identification of degraded forensic samples. Previous biomedical research also showed that microRNAs are tissue-specific, indicating the cell lines contained within these tissues can be separated by unique sequences of microRNAs, referred to as biomarkers. Unlike protein-based tests, microRNAs can be amplified using polymerase chain reaction (PCR) hundreds of times to remove issues with not having enough material present.
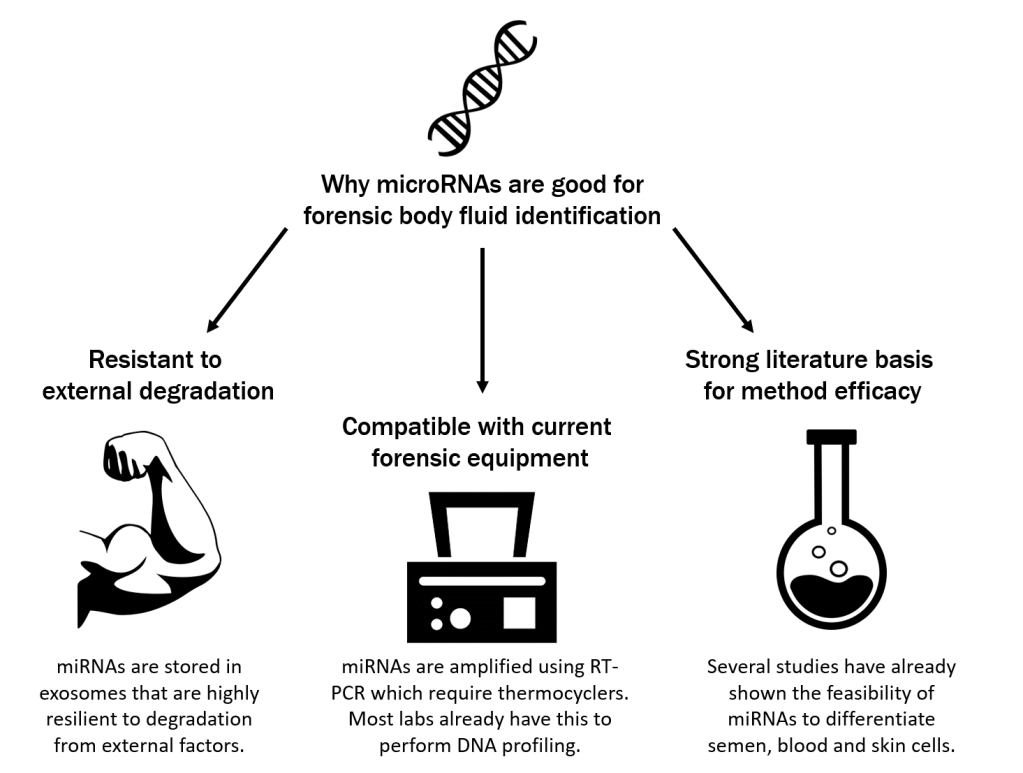
Previous groups have been able to separate semen, blood and skin cells based on the unique expression of microRNAs – but not, as Tamaki and colleagues point out, saliva and vaginal fluids, which contain similar unique microRNA biomarkers. Tamaki group’s did not propose a new experimental method, but a new a statistical model, to separate blood, semen, saliva and vaginal fluids. Statistical models allow scientists to interpret their data in different ways to draw out the most information. The study’s model, called probabilistic discrimination, creates the maximum “distance” between groups based on a principal component (specific microRNAs) that creates new groupings (explained later, called PLS components) that sharpen group distinction (such as body fluids).
After extracting the microRNAs and amplifying the sequence using reverse-transcriptase PCR (they need to copy or transcribe the sequence from RNA to DNA first to amplify the sequence, hence the name reverse-transcriptase), the scientists listed the unique microRNA biomarkers found in each fluid. Then, they sorted the unique microRNAs into groups called partial least squares (PLS) components that allowed the most “distance” between each body fluid. By fitting their groupings of data into three PLS groups and plotting each PLS component on an axis, first author Fujimoto and colleagues created a model that taught the software how to distinguish between blood, semen, saliva and vaginal fluid with low error.
Much in a way an athlete trains on a practice course before competing on the actual stage, statisticians “train” their programs on a practice set (known as the “training set” and shown as circles in the 3D plot) before evaluating their model on the actual data (known as the “testing set” and shown as crosses in the 3D plot). This training-and-testing method allows scientists to see how well their model predicts the answer (in this case, the body fluid) from the given information. Fujimoto’s model correctly identified which samples were blood, semen, vaginal fluid or saliva based on the input miRNA profile information. However, there was some overlap between the saliva and vaginal fluid samples that could not be fixed by changing the model [Fig 2].
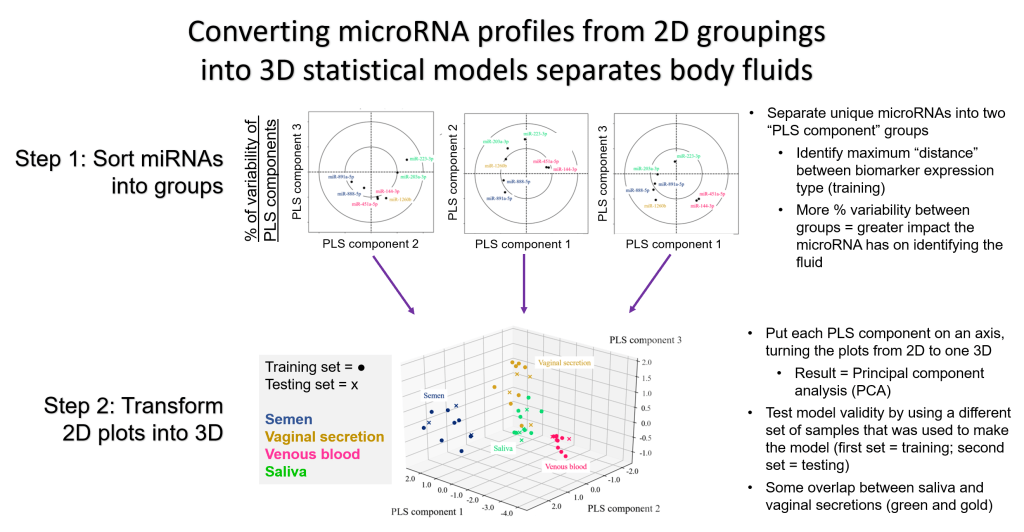
Fujimoto et al. also considered two unique factors that impact semen and vaginal fluid; infertility and menstruation. Male infertility influences the quantity and quality of sperm in semen, along with other chemical and protein levels such as fructose, testosterone and follicle-growth hormone. Women at different stages of the menstrual cycle have different levels of vaginal discharge, bacterial species and estrogen. Tamaki’s team wondered if either of these conditions impact the microRNA expression in semenial or vaginal body fluids. By monitoring their two semen-specific and seven vaginal fluid identified microRNA strands, they found no difference for either condition, including with azoospermatozoic (no sperm) or pre-/post-ovulation.
Though the study did not include samples of artificially degraded body fluids or casework examples, the work of Fujimoto and colleagues in Tamaki’s lab adds to a growing body of evidence that microRNA profiles, coupled with statistical analysis, can identify and distinguish between different body fluids. As the process of obtaining microRNAs requires equipment already present in forensic laboratories, I anticipate commercial entities need only to create RT-PCR kits using ingredients for forensic body fluids with rigorous quality control to see this method become a standard of the forensic field.
| Title | Distinct spectrum of microRNA expression in forensically relevant body fluids and probabilistic discriminant approach |
| Authors | Shuntaro Fujimoto, Sho Manabe, Chie Morimoto, Munetaka Ozeki, Yuya Hamano, Eriko Hirai, Hirokazu Kotani & Keiji Tamaki |
| Journal | Nature Scientific Reports |
| Year | 2019 |
| Link | https://www.nature.com/articles/s41598-019-50796-8 |
